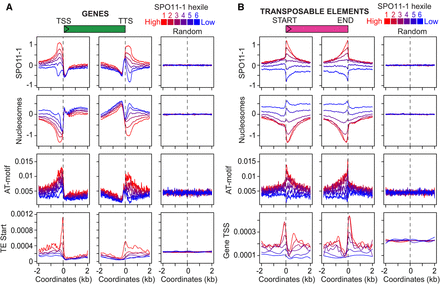
Nucleosomes, AT-sequence motifs, and SPO11-1-oligonucleotides within genes and transposons. (A) Density of SPO11-1-oligos (Z-score standardized log2[SPO11-1-oligos/gDNA]), nucleosome occupancy (Z-score standardized log2[MNase/gDNA]), crossover-enriched AT-motifs (Choi et al. 2013), and TE start coordinates, in 4-kb windows around gene TSSs or TTSs, or the same number of random positions. Genes are grouped according to SPO11-1-oligo promoter or terminator hexile: (red) highest; (blue) lowest. (B) As for A but analyzing transposon SPO11-1-oligo hexiles and showing gene TSS density.











