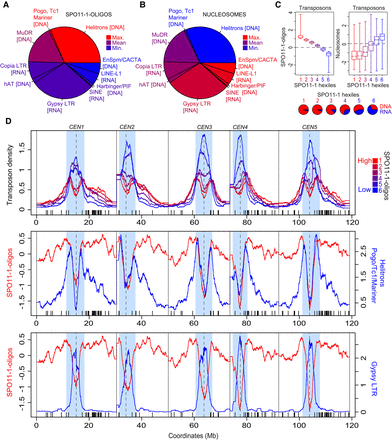
Meiotic recombination and chromatin variation between A. thaliana transposons. (A) Pie chart showing A. thaliana transposon families, with slice size proportional to physical length, and color-coded according to SPO11-1-oligo (Z-score standardized log2[SPO11-1-oligos/gDNA]) levels. Inset colored boxes are shaded equivalent to the genome-wide mean (purple), minimum (blue), and maximum (red) values. (B) As for A, but analyzing nucleosome occupancy (Z-score standardized log2[MNase/gDNA]). (C) Box plots showing SPO11-1-oligo and nucleosome occupancy, according to transposon SPO11-1-oligo hexile groups, with horizontal lines indicating the genome average value. Inset pie charts show the proportion of DNA (red) and RNA (blue) transposons for each SPO11-1-oligo transposon hexile. (D) Density of transposons through the A. thaliana genome according to SPO11-1-oligo hexile: (red) highest; (blue) lowest. x-Axis ticks indicate NBS-LRR gene homolog positions. Plotted beneath are SPO11-1-oligos (red) versus the density of Helitron/Pogo/Tc1/Mariner class DNA transposons (blue), or Gypsy RNA transposons (blue).











