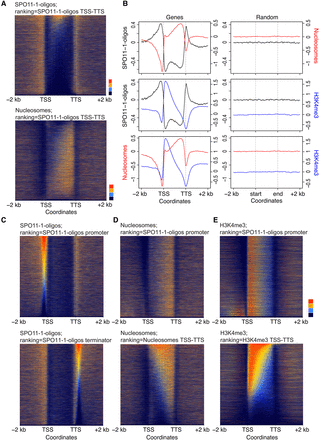
SPO11-1-oligonucleotide enrichment in gene promoter and terminator nucleosome-free regions. (A) Heat maps of SPO11-1-oligos (upper) and nucleosome occupancy (lower) within gene transcriptional units (between transcriptional start [TSS] and termination [TTS] sites) and 2-kb flanking regions. The rows represent individual genes, which have been ordered by descending SPO11-1-oligo values between TSS and TTS. For shading, SPO11-1-oligo and nucleosome values equal to defined quantiles were mapped linearly to a vector of six colors: dark blue (lowest), blue, light blue, yellow, orange, red (highest). (B) Density of SPO11-1-oligos (black; log2[SPO11-1-oligos/gDNA]), nucleosome occupancy (red, nucleosomes; log2[MNase/gDNA]), and H3K4me3 (blue; log2[ChIP/input]) in wild type, across gene transcriptional units (TSS to TTS), and in flanking 2-kb windows. The right-hand column shows results from analysis of the same number of randomly chosen positions. (C) Heat maps as for A, but showing SPO11-1-oligos around genes ordered by descending SPO11-1-oligo levels in gene promoters (upper, 500 bp upstream of TSS) or gene terminators (lower, 500 bp downstream from TTS). (D) Heat maps as for A, but showing nucleosomes around genes ordered by descending SPO11-1-oligo levels in gene promoters (upper), or by descending nucleosome levels within TSS–TTS (lower). (E) Heat maps as for A, but showing H3K4me3 around genes ordered by descending SPO11-1-oligo levels in gene promoters (upper) or by descending H3K4me3 levels within TSS–TTS (lower).











