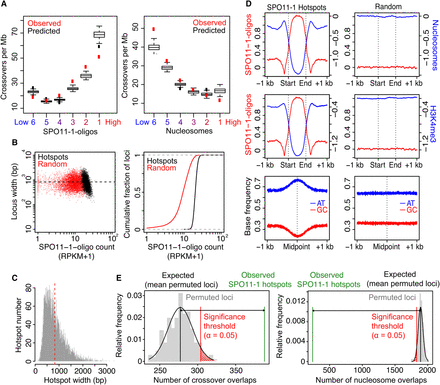
A. thaliana SPO11-1-oligonucleotide hotspots and crossovers. (A) Plots showing observed (red) overlap of SNP intervals with crossovers per megabase. SNP intervals (n = 479,888) were divided into six groups (hexiles) following ranking by mean SPO11-1-oligo levels (log2[SPO11-1-oligos/gDNA]; group 1 = highest, group 6 = lowest). Data were modeled with the glm2 function in R, using the binomial family with logistic link function and the formula: CO ∼ SPO11-1-oligos + nucleosomes + H3K4me3 + DNA methylation + width + interactions. Using this logistic model, we then obtained the probability of windows within each hexile overlapping a crossover (black box plots). (B) SPO11-1-oligo counts per hotspot (RPKM + 1) plotted against hotspot widths (bp; black) compared with equivalent analysis of random loci of the same number and widths (red). Also shown is a cumulative distribution curve plotting hotspots (black) and random loci (red), according to ranked SPO11-1-oligo counts (RPKM + 1). (C) Histogram of ranger-defined SPO11-1-oligo hotspot widths (bp), with the mean indicated by the red vertical dashed line. (D) Average profiles of SPO11-1-oligos (red, Z-score standardized log2[SPO11-1-oligos/gDNA]), nucleosomes (blue, Z-score standardized log2[MNase/gDNA]), and H3K4me3 (blue, Z-score standardized log2[ChIP/Input]) within SPO11-1-oligo hotspots, scaled to a common width, and in 1-kb flanking windows (left). Equivalent randomly positioned loci were analyzed in the same way (right). Also shown are plots analyzing the relative frequency of AT (blue) and GC (red) bases around SPO11-1-oligo hotspots or random loci. (E) Histograms show the relative frequency distribution of 10,000 sets of permuted (randomly positioned) loci, which are of the same number and widths as the SPO11-1-oligo hotspots, with regard to the number of permuted loci that overlap one or more crossovers (left) or highly positioned nucleosomes (right; gray bars). The number of SPO11-1-oligo hotspots observed (green vertical line) or expected (black vertical line; mean overlaps for permuted loci) to overlap one or more loci, together with the significance threshold (red vertical line; α = 0.05) for a difference between observed and expected overlaps, are indicated.











