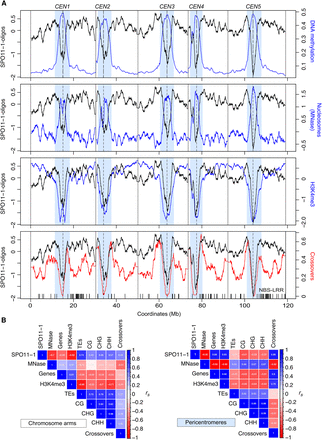
Genomic landscape of SPO11-1-oligonucleotides, crossovers, euchromatin, and heterochromatin. (A) SPO11-1-oligos (black, Z-score standardized log2[SPO11-1-oligos/gDNA]), nucleosome occupancy (blue, Z-score standardized log2[MNase/gDNA]) (Choi et al. 2016), and H3K4me3 (blue, Z-score standardized log2[ChIP/input]) levels were calculated in adjacent 10-kb windows and plotted along the A. thaliana chromosomes, using a rolling average. DNA methylation (blue, proportion of methylation in all sequence contexts) (Stroud et al. 2013) and crossovers (red) (Choi et al. 2016; Serra et al. 2018) were analyzed in the same way using 200-kb and 10-kb windows, respectively. The centromeric assembly gaps are indicated by vertical dashed lines, and telomere positions are indicated by vertical solid lines. The pericentromeres are shaded light blue and are defined as regions surrounding the centromere with greater than average DNA methylation. x-Axis ticks indicate the positions of NBS-LRR gene homologs (Choi et al. 2016). (B) Matrices showing Spearman's rank correlation coefficient between the listed parameters, with 10-kb windows used for correlations and shading proportional to the value. Matrices were calculated separately for the chromosome arms and pericentromeres.











