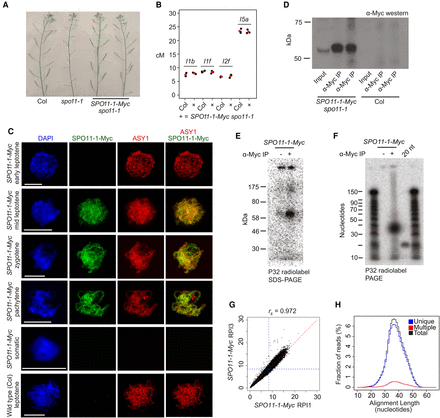
Purification and sequencing of A. thaliana SPO11-1-oligonucleotides. (A) Inflorescences from wild type (Col), spo11-1, and SPO11-1-Myc spo11-1. Greater silique length indicates higher fertility. (B) Crossover frequency (cM) measured using fluorescent crossover reporter lines (FTLs I1b, I1f, I2f, and I5a) in wild type (Col) or SPO11-1-Myc spo11-1 (+). Mean values are shown in red. (C) Nuclei from SPO11-1-Myc or wild type (Col) immunostained using α-Myc (green) or α-ASY1 (red) antibodies and stained for DAPI (blue). Scale bars, 10 μM. (D) α-Myc Western blot from SPO11-1-Myc or Col extracts, before and after α-Myc immunoprecipitation (α-Myc-IP). Biological replicate samples are shown. (E) Detection of end-radiolabeled SPO11-1-Myc complexes following immunoprecipitation and SDS polyacrylamide gel electrophoresis (SDS-PAGE). (F) Detection of end-radiolabeled, purified SPO11-1-oligos following proteinase K digestion of immunoprecipitates and polyacrylamide gel electrophoresis (PAGE). A labeled 20-base oligonucleotide (20 nt) was run alongside as a size control. (G) Scatter plot showing correlation of library size normalized SPO11-1-oligo values calculated in adjacent 10-kb windows between wild-type libraries RPI1 and RPI3. Blue dotted lines indicate genome average values. Spearman's rank correlation coefficient (rs) is printed above the plot. (H) Histogram showing lengths of uniquely aligning (blue), multiply aligning (red), and total (black) SPO11-1 reads from library RPI48.











