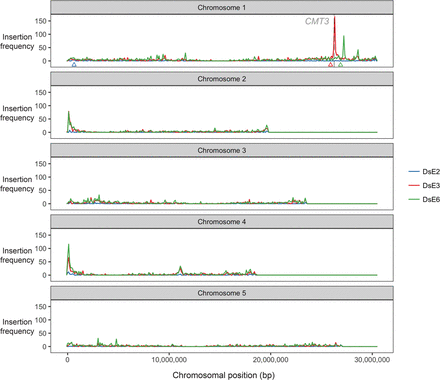
Transposon insertions in CMT3 appear to induce recombination. Enhancer trap DNA transposons (DsE) based on the maize transposable element Dissociation (Ds) were introduced into the A. thaliana genome on a T-DNA with a negative selectable marker (Sundaresan et al. 1995). Three independent DsE launch-pads were mapped to Chromosome 1: DsE2 (633,819 bp, blue triangle), DsE3 (25,893,665 bp, red triangle), and DsE6 (26,877,449 bp, green triangle). After introducing Activator transposase, 9622 transpositions of DsE were isolated in F2 progeny by selecting for the transposon, but against the launch-pad, to select against linked transpositions which would otherwise be highly favored. New DsE insertions were mapped by sequencing flanking DNA. Transpositions from each Chromosome 1 launch-pad are displayed in 100-kb bins, and most were unlinked, including two hotspots corresponding to nucleolar organizer regions at the ends of Chromosomes 2 and 4. However, two much sharper insertions hotspots, on Chromosome 1, were specific for closely linked launch-pads, DsE3 (red triangle) and DsE6 (green triangle), respectively. The hotspot immediately distal to DsE3 (∼4% of transpositions launched from DsE3) was in and immediately around the CMT3 gene (26,248,318–26,253,585 bp).











