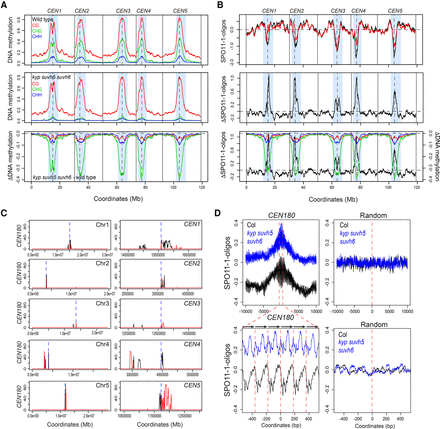
Elevated SPO11-1-oligonucleotides levels in centromeres of kyp suvh5 suvh6 H3K9me2 mutants. (A) Plots of the A. thaliana chromosomes on a continuous x-axis are shown. Analysis of DNA methylation frequency in CG (red), CHG (green), and CHH (blue) from published data in wild type (Col) (top) or kyp suvh5 suvh6 (middle) (Stroud et al. 2013). A kyp suvh5 suvh6 minus wild type differential (ΔDNA methylation) plot is also shown (bottom). Vertical dotted lines indicate the position of the centromere assembly gaps and vertical solid lines indicate telomeres. The pericentromeres, defined by higher than average DNA methylation, are indicated by light blue shading. (B) Plots showing log2(SPO11-1-oligos/gDNA) (SPO11-1-oligos) in wild type (Col, black) and kyp suvh5 suvh6 (red) (top), in addition to the kyp suvh5 suvh6-wild type differential (ΔSPO11-1-oligos) (middle), which is also overlaid with the ΔDNA methylation from A (bottom). Plots are annotated as in A. (C) Plots of the A. thaliana chromosomes showing the density of CEN180 repeats on forward (black) and reverse (red) strands. The TAIR10 centromeric assembly gaps are shown by the dotted blue line. The plots on the left show whole chromosomes, whereas the plots on the right show a close-up of the regions surrounding the centromere assembly gaps. (D) The density of log2(SPO11-1-oligos/gDNA) (SPO11-1-oligos) was analyzed in ±10-kb windows surrounding matches to the CEN180 consensus in wild type (black) or kyp suvh5 suvh6 (blue). The lower plot corresponds to a blow-up of a 1-kb window around the center of the upper plot. The lower plot highlights the approximate positions of CEN180 repeats using arrows and red dotted lines. Also shown is an identical analysis performed for the same number of randomly chosen sites.











