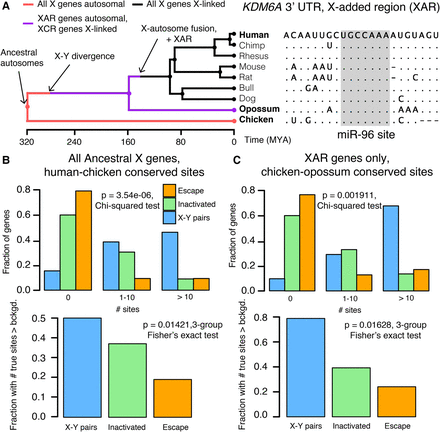
Heterogeneities in X-linked miRNA targeting were present on the ancestral autosomes. (A) Example reconstruction of an ancestral miR-96 target site in the 3′ UTR of KDM6A, an X-linked gene in the X-added region (XAR) with a surviving Y homolog. (Left) Species tree overlaid with events in mammalian sex chromosome evolution. (Right) Multiple sequence alignment; dots in nonhuman species indicate identity with human sequence; dashes indicate gaps in alignment. (B) Distributions of sites conserved between 3′ UTRs of human and chicken orthologs (top) or comparisons to background expectation (bottom; see Methods) for human X-Y pairs (n = 16), X-inactivated genes (n = 251), and X escape genes (n = 42). (C) Statistics as in B, but using sites conserved between chicken and opossum 3′ UTRs only for genes in the XAR: X-Y pairs (n = 11), X-inactivated genes (n = 58), and X escape genes (n = 27).











