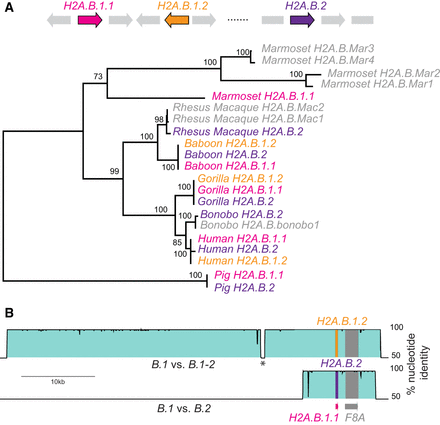
Concerted evolution by gene conversion among primate H2A.B genes. (A) Maximum-likelihood nucleotide phylogeny of representative primate H2A.B genes rooted using pig as an outgroup. Bootstrap values are shown for nodes with >50% support. A cartoon shows the loci containing ancestral duplications H2A.B.1.1 (pink), H2A.B.1.2 (orange), and H2A.B.2 (purple), and gene names are colored accordingly in the tree. We also include some genes that are either in nonsyntenic loci (representing additional duplicates) or were on short scaffolds so could not be assigned to a locus (four from marmoset, two from rhesus macaque, and one from bonobo, denoted .Mar1 to .Mar4; .Mac1, .Mac2; and .bonobo1, respectively). (B) VISTA plots showing very high nucleotide identity across large genomic regions flanking the three human H2A.B genes (proximal to F8A), comparing H2A.B.1.1 genomic sequence as a reference to H2A.B.1.2 (upper plot) and H2A.B.2 (lower plot). An interruption in high identity (asterisk) is due to a recent transposon insertion.











