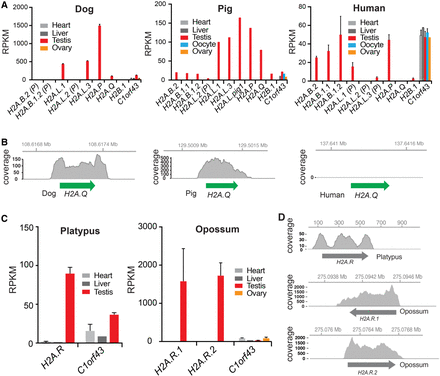
Expression of short histone variants. (A) Analyses of short H2A expression using public RNA-seq data from selected tissues of dog, pig, and human. Error bars show standard deviation of biological replicates, where available. Expression of the germ cell–specific H2B.1 variant and a housekeeping gene C1orf43 (Eisenberg and Levanon 2013) are shown for comparison (more controls are shown in Supplemental Fig. S3). Mapped reads are shown in reads per kilobase per million mapped (RPKM). (B) RNA-seq coverage over the H2A.Q locus in dog, pig, and human. Coordinates along the assembled X Chromosome are indicated at the top. (C) RNA-seq analyses of platypus and opossum H2A.Rs, and one housekeeping gene (as in A), across somatic and reproductive tissues. Error bars as in A. (D) RNA-seq coverage as in B. Coordinates along the annotated contig or chromosome (Chr 6 for opossum H2A.Rs and contig111576 for platypus) are shown at the top.











