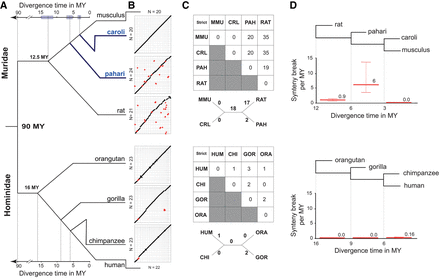
Muridae genomes undergo large chromosomal rearrangements in punctuate bursts, resulting in greater structural diversity than primates. (A) Phylogenetic tree showing that the divergence time of the four Muridae species mirrors that of the four Hominidae species. The Mus species in blue were sequenced and assembled for this study. The 95% confidence interval of the divergence time estimation is shown by the shaded boxes (Supplemental Methods SM1.16). (B) Dot plots of whole-genome pairwise comparison between Mus musculus and the three other Muridae (top), and between human and the three other Hominidae (bottom). The chromosomes of Mus musculus and human were ordered by chromosome number. The chromosomes of the other species were ordered to optimize the contiguity across the diagonal. Red dots represent large (>3 Mb) inter-chromosomal rearrangements (fusion/fission and translocation). (C) Matrix of neighbor-joining tree of synteny breaks involving inter-chromosomal rearrangement for Muridae and Hominidae: (MMU) Mus musculus; (CAR) Mus caroli; (PAH) Mus pahari; (RAT) rat; (HUM) Human; (CHI) chimpanzee; (GOR) gorilla; (ORA) orangutan. (D) The rate of synteny breaks between sequential internal branch points of the Muridae and Hominidae clades. Muridae have undergone a punctuate increase in the rate of syntenic breaks between 3 and 6 MYA.











