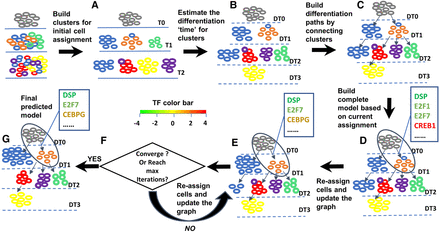
Learning differentiation models from single-cell RNA-seq data. (A) Initial clusters are determined using spectral clustering. T0, T1, and T2 represent the measurement time. (B) Initial “differentiation time” is estimated for clusters based on difference with clusters for the first time point. DT0, DT1, DT2, and DT3 denote the estimated differentiation time. (C) Differentiation paths are constructed by connecting clusters at lower levels to their most similar parent at the level above them. (D) Regulating TFs are determined for each edge. TFs are colored based on their expression change along the edge. (Red) Increased expression; (green) decreased expression; (blue) stable expression. Shades represent the extent of the expression change. (E) Initial model. (F) Iterating between cells and state reassignments and parameter learning until convergence. (G) Final model.











