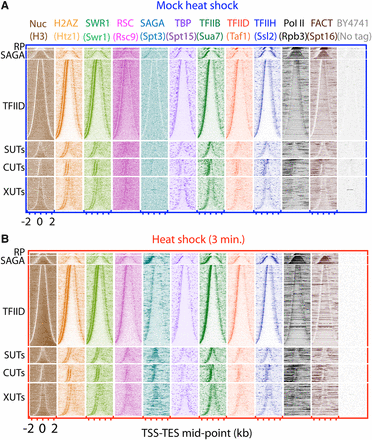
Factor occupancy at all genes before and after heat shock. (A) Yeast cultures were either mock-treated (upper set of panels) or (B) heat shocked (lower) for 3 min at 37°C. ChIP-exo tag 5′ ends were plotted relative to transcription unit midpoints (start to end from left to right). Rows are sorted by unit length and grouped by class: ribosomal protein genes (RP, n = 130), SAGA-dominated genes (SAGA, n = 451), TFIID-dominated genes (TFIID, n = 4260), and noncoding SUTs (n = 846), CUTs (n = 924), and XUTs (n = 1720). All rows across data sets are linked.











