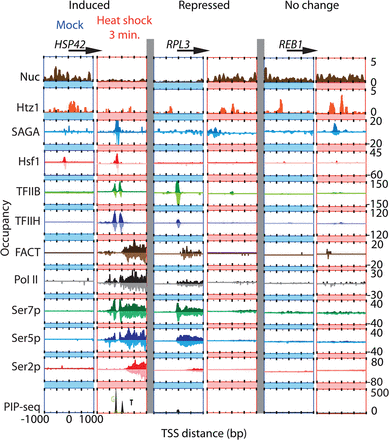
Factor occupancy at three genes before and after heat shock. Smoothed distribution of unshifted ChIP-exo tag 5′ ends (exonuclease stop sites) on forward and reverse strands (with the latter being inverted) on induced HSP42, repressed RPL3, and housekeeping REB1 genes upon mock or after 3 min of heat shock. Nucleosomes (Nuc) represent fragment midpoints of MNase-digested H3-immunoprecipitated chromatin. All TSSs are oriented such that the direction of transcription is to the right. The bottom panel of each column corresponds to melted PIC DNA derived from TFIIB PIP-seq. Foreground “T” nucleotides are in black and background “G” nucleotides are in green. Y-axes are scaled for each factor to fit the frame and are all on the same scale for a particular factor (row).











