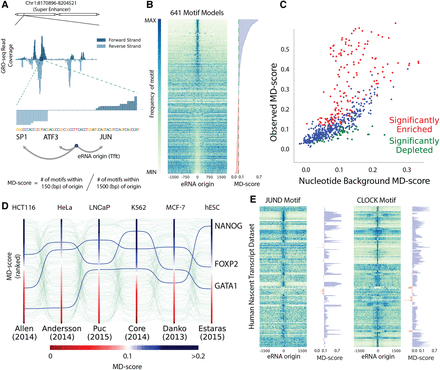
Motif colocalization with eRNA origins varies by cell type. (A) An example locus of GRO-seq, the inferred eRNA origin, and computation of “motif displacement” (MD) and the associated MD-score. (B) Each row is a TF motif model, and each column is a bin of a histogram (100) where heat is proportional to the frequency of a motif instance at that distance from an eRNA origin. (C) A comparison between the expected MD-score for a motif model (x-axis) and the observed MD-score in a K562 GRO-cap experiment (Core et al. 2014). Red and green dots indicate a P-value <10−6 above or below expectation hypothesis tests, respectively. (D) MD-scores were computed and ranked under six nascent transcription data sets. (E) Each row corresponds to a nascent data set, and each column relates to motif frequency. These MD distributions are shown for two demonstrative examples (JUND and CLOCK) and the associated MD-scores, sorted by publication.











