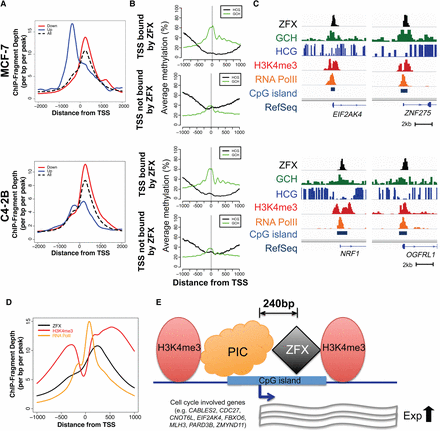
The relationship between ZFX binding and chromatin structure at promoters. (A) Average ZFX ChIP-seq signals ±2 kb from the TSS of down-regulated (red), up-regulated (blue), and all (black) genes bound by ZFX in MCF-7 (top) and C4-2B (bottom). (B) Endogenous DNA methylation (HCG) (black) and the accessibility (GCH) (green) levels from NOMe-seq data ±1 kb from the TSS of active promoters bound by ZFX and from the TSS of active promoters not bound by ZFX in MCF-7 (top) and C4-2B (bottom). (C) Examples of ZFX binding sites with ZFX ChIP-seq, NOMe-seq, H3K4me3 ChIP-seq, and RNA Polymerase II ChIP-seq signals in MCF-7 (top) and C4-2B (bottom). (D) Average ZFX (black), H3K4me3 (red), and RNA Polymerase II (orange) ChIP-seq signals ±1 kb from the TSS of genes bound by ZFX in MCF-7. (E) A model demonstrating the relationship of ZFX to other components of CpG island promoter structure. ZFX binds at +240 bp in the nucleosome-depleted region of CpG island promoters, between the general transcription preinitiation complex (PIC) and the first nucleosome in the transcribed region. When ZFX is bound to this downstream site, it increases the expression levels of genes involved in cell proliferation; the wavy lines represent RNA levels.











