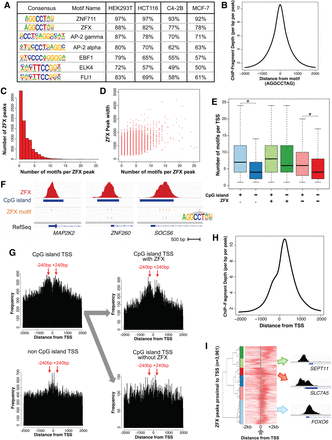
Characterization of ZFX motifs and binding sites. (A) Motifs enriched at summits of the ZFX binding sites and the percentage of ZFX peaks having each motif for HEK293T kidney, HCT116 colon, C4-2B prostate, and MCF-7 breast cancer cells; the same motifs were identified in the set of ZFX peaks located near a TSS and the set of distal ZFX peaks. (B) Average ZFX ChIP-seq signal in C4-2B relative to ±2 kb from the motif AGGCCTAG. (C) Histogram of the number of motifs per ZFX peak. (D) Scatterplot of the relationship between the number of motifs per ZFX peak and ZFX peak width. (E) Number of AGGCCTAG motifs per TSS region (±2 kb from the TSS) for different groups of promoters: (light blue) CpG island promoters; (blue) non-CpG island promoters; (light green) CpG island promoters bound by ZFX; (green) non-CpG island promoters bound by ZFX; (light pink) CpG island promoters not bound by ZFX; (red) non-CpG island promoters not bound by ZFX. Comparisons of data sets that show a significant difference (adjusted P-value <0.05) are marked with an asterisk. (F) Example of ZFX binding site and motif position at CpG island promoters having one (MAP2K2) or more (ZNF260, SOCS6) motifs. (G) Frequency of AGGCCTAG motifs located ±2 kb from the TSS of CpG island promoters, of non-CpG island promoters, of CpG island promoters bound by ZFX, and of CpG island promoters not bound by ZFX; a comparison to results obtained using a scrambled motif can be found in Supplemental Figure S2. (H) Average ZFX ChIP-seq signal ±2 kb from the TSS of promoters bound by ZFX in MCF-7. (I) Heatmap representing unsupervised clustering of ZFX ChIP-seq signals in MCF-7 cells at promoters bound by ZFX that only have one TSS in a ±2 kb region (n = 3961). Also shown are examples of ZFX binding sites from the light green cluster (promoters having a ZFX peak only upstream of the TSS), the red cluster (promoters having a ZFX peak both up and downstream from the TSS), and the light blue cluster (promoters having a ZFX peak only downstream from the TSS). Genes from each cluster are listed in Supplemental Table S3F.











