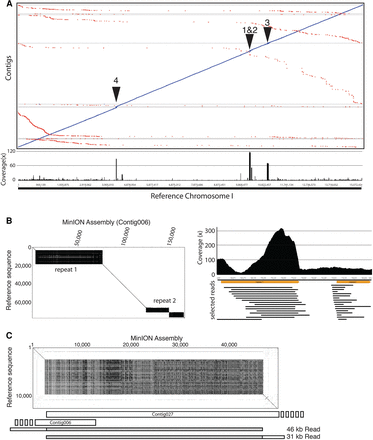
MinION sequencing and assembly reveals genome expansion of C. elegans repeat regions. (A) LASTZ alignment of contigs to the C. elegans reference Chromosome I. Below is a plot of coverage of the ‘All data’ assembly contig sequences against the reference. Note the high coverage areas (indicated as 1–4) that correlate with repeat regions. (B) (Left) Dot plot of the MinION-derived contig containing repeats 1 and 2 against the reference sequence. (Right) Plot of sequence read coverage (reads > 5 kb) in the repeat regions. Selected reads spanning the expanded repeats are shown below. Note the high coverage of repeat 1, which suggests that this repeat is even larger than predicted in the contig shown. Repeat 2 read coverage, which is similar to the average read coverage across the genome, suggests that this repeat expansion is correct. (C) Dot plot of the end termini of contig006 and contig027 against the reference sequence demonstrating the expansion of repeat 3. Two of the longest reads mapping to this repeat are shown below with the repeat highlighted in gray.











