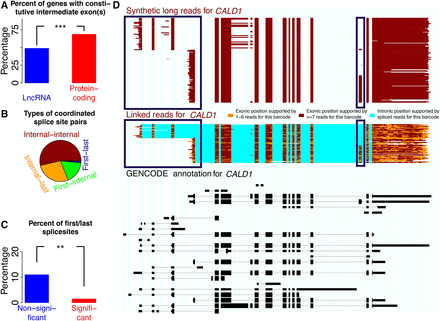
Coordination between first alternative donors and last alternative acceptors. (A) Percent of exon pairs that are always separated by at least one intermediate exon for lincRNAs and for protein-coding genes. (B) Frequency among all coordinated pairs of pairs of internal splice sites (“Internal-internal”), pairs of an internal and a last splice site (“Internal-last”), pairs of a first splice site and an internal exon (“First-internal”), and pairs of a first and a last splice site (“First-last”). (C) Percentage of pairs of a first and a last splice site among coordinated (FDR < 0.05) and noncoordinated pairs. (D) Bottom, black track: GENCODE annotation. Middle, colored track: spISO-seq data, with each line representing one molecule. Top, red-brown track: SLR-RNA-seq data with each line representing one molecule. Blue boxes highlight first exon and TSS choice (left blue boxes) and internal exon inclusion (right blue boxes). Inclusion of the alternative internal exon occurs only when the downstream first exon/TSS is chosen.











