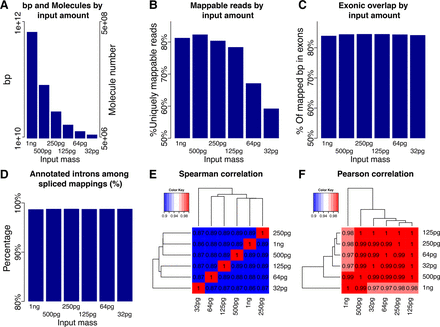
Exploration of low input capacities using shallow sequencing of a MiSeq. (A) Number of molecules and bases for different input amounts. (B) Percentage of reads that were uniquely mappable for all input amounts. (C) Percentage of mapped bases that fall onto annotated GENCODE exons for all input amounts. (D) Percentage of annotated introns among all spliced mappings for all input amounts. (E) Heat map of pairwise Spearman correlations of FPKMs for all input amounts. (F) Heat map of pairwise Pearson correlations of FPKMs for all input amounts.











