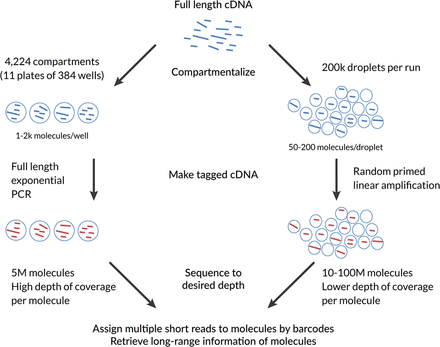
Comparative outline of the spISO-seq and the previously published SLR-RNA-seq approach. (1) Both approaches rely on the principle of compartmentalization. The fewer cDNA molecules are separated into one compartment, the lower the probability of having two nonidentical molecules from the same gene. SLR-RNA-seq employs 1000–2000 molecules per well on 384-well plates, while spISO-seq employs 50–200 molecules per droplet for a total of ∼200,000 droplets. (2) SLR-RNA-seq performs a full-length PCR that is exponentially amplifying all molecules in a well, while spISO-seq performs a linear randomly primed amplification. (3) In both approaches, the amplified product is short-read-sequenced using barcodes that identify the compartment (well or droplet of origin) and (based on that, most of the time only one molecule per gene is observed per compartment) the molecule of origin. (4) All short reads originating from the same molecule of origin are then collectively analyzed to retrieve long-range information within molecules.











