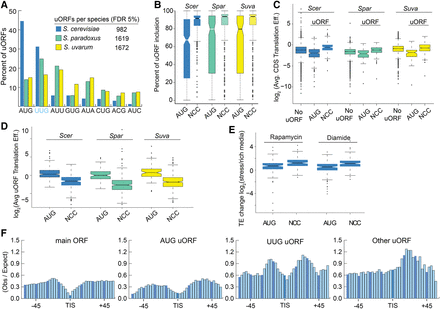
Comparing AUG and NCC uORFs in Saccharomyces yeasts. (A) In S. cerevisiae, nearly 45% of uORFs initiate with AUG, compared with ∼15% in S. paradoxus and S. uvarum. (B) AUG uORFs had significantly lower rates of transcript inclusion (the fraction of a gene's transcripts that contain the uORF; AUG median 73%) compared to NCCs (92%), suggesting alternative transcription initiation plays a greater role in AUG uORF regulation than NCC type. (C) Genes with AUG uORFs have lower translation efficiencies, whereas genes with NCC uORFs have higher translation efficiencies than genes without uORFs. (D) AUG uORFs have higher translation efficiencies than NCC uORFs. (E) The translation efficiencies of AUG and NCC uORFs increase with stress, and NCC uORFs increase more. (F) A metagene analysis of RBP interactions around main ORF and uORF start codons is shown using public data (Freeberg et al. 2013). AUG and UUG uORFs have similar patterns of protein occupancy with peaks of occupancy upstream of and downstream from the start codon. This pattern was not observed for other NCC uORFs, which only exhibited a significant peak downstream from the TIS.











