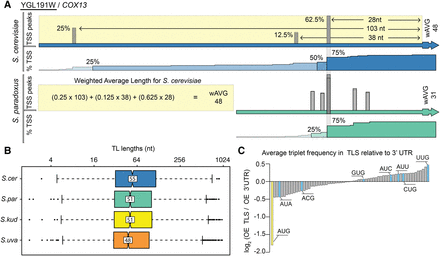
Transcript leader evolution in Saccharomyces sensu stricto. (A) An example comparison of TLs of COX13 (YGL191W) from S. cerevisiae and S. paradoxus is shown. Although the most frequent TSS has been conserved (28 nt TL), the most upstream TSS has diverged. The weighted average (wAVG) length for each transcript leader was calculated to compare transcript leader lengths among species. (B) S. cerevisiae has significantly longer TLs than the other species. (C) The observed/expected ratio (OE) of each 3-nt triplet in 5′ UTRs versus 3′ UTRs is plotted. Relative to the 3′ UTR, the TLSs are de-enriched in AUG triplet nucleotides (black), and enriched in five of seven near-cognate codon (NCC) triplets (gray). The data presented are an average of values from the four species; individual values are plotted in Supplemental Figure S14.











