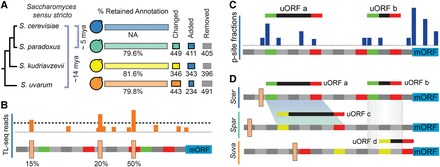
Overview. (A) The genomes of nonmodel species were reannotated to include ncRNA and correct previous errors, resulting in changes to ∼20% of genes on average (Methods). (B) Transcript leaders and polyadenylation sites were annotated for four Saccharomyces sensu stricto species using TL-seq (Arribere and Gilbert 2013) and pA-seq (Yoon and Brem 2010). TL-seq and pA-seq peaks were trend filtered for FDR control (β = 0.001) and evaluated for their usage frequency. (C) Ribosome profiling data from S. cerevisiae, S. paradoxus, and S. uvarum were analyzed to quantify the locations of ribosome P-site occupancy (dark blue peaks). These results were used to identify candidate uORFs. (D) Candidate uORFs were evaluated and scored using the uORF-seqr machine learning algorithm. uORFs with significant scores were retained for further analysis. The transcript leaders of gene homologs were aligned to identify uORF homologs (marked by shaded maps). These were identified by both local sequence alignment (uORFs a and c) and by positional overlap (uORFs b and d).











