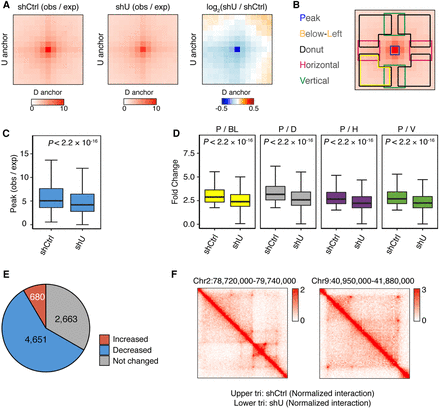
The intensity of chromatin loops decreases upon HNRNPU depletion. (A) Mean aggregate signals of 7994 loops by summing submatrices surrounding each peak (10 kb × 10 kb for each pixel). (B) Schematic view of loop center (peak) and adjacent areas: below-left (yellow), donut (black), horizontal (purple), and vertical (green). (C) Box plots showing the intensity of peaks in shCtrl- and shU-treated cells. P-values were obtained by Wilcoxon rank-sum test. (D) Box plots showing fold change of peak signal relative to adjacent areas. P-values were obtained by Wilcoxon rank-sum test. (E) Percentage of changed chromatin loops upon HNRNPU depletion. (F) Representative regions showing decreases of chromatin loops.











