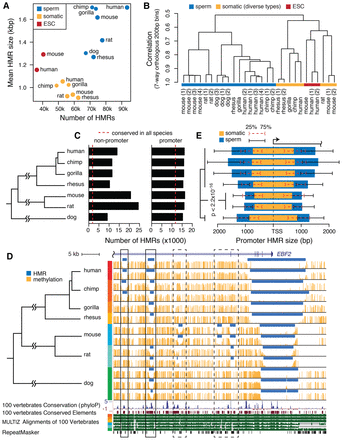
Mammalian sperm methylome characteristics. (A) The number and average size of HMRs in native assemblies. (B) Hierarchical clustering of aligned seven-way orthologous methylomes of multiple species and cell types. (C) The number of promoter HMRs and nonpromoter HMRs in seven-way orthologous sperm methylomes. Dashed lines indicate the average number of conserved HMRs across species. (D) Sperm DNA methylation of seven species in an example orthologous region. Methylome alignment is shown along with conservation tracks from MULTIZ alignment of 100 vertebrates and human repeat elements by RepeatMasker (Smit et al. 2013–2015). Regions in solid boxes show divergent methylation states at well-conserved elements. See zoomed-in browser image for dashed boxes in Supplemental Figure S2A. (E) The median size of hypomethylated regions upstream of and downstream from TSS in somatic (orange) and sperm (blue) methylomes of different species. HMR sizes are measured in their respective native genomes. Whiskers mark the 25th and 75th percentiles of HMR sizes upstream of or downstream from TSS. Wilcoxon rank-sum tests for all pairs of species between rodent species and nonrodent species showed significantly narrower HMRs in rodents.











