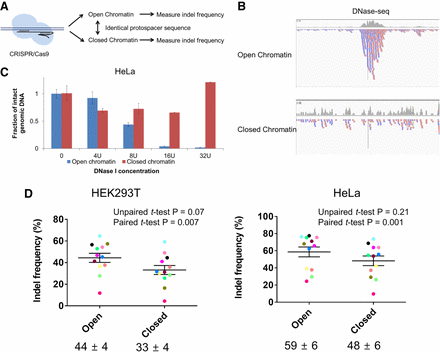
Effects of chromatin structure on Cas9 editing efficiency. (A) Schematic overview of the method for investigating the effects of chromatin structure on Cas9 editing efficiency. (B) Representative IGV images obtained using ENCODE DNase-seq data at two on-target sites with the same Cas9 target sequence present in both open and closed chromatin regions. (C) Relative fractions of intact genomic DNA not cleaved by DNase I measured using real-time quantitative PCR at the Cas9 target site in HeLa cells. The range 0–32U denotes the concentration of DNase I. Error bars, SEM from at least three independent experiments. (D) SpCas9-induced mutation frequencies at 12 pairs of endogenous target sites with the same DNA sequence present in both open and closed chromatin regions in HEK 293T or HeLa cells. Indel frequencies were measured using targeted deep sequencing. Pairs of Cas9 target sites are represented with dots with different colors. Mean indel frequencies ± SEM are shown (n = 12 target sites).











