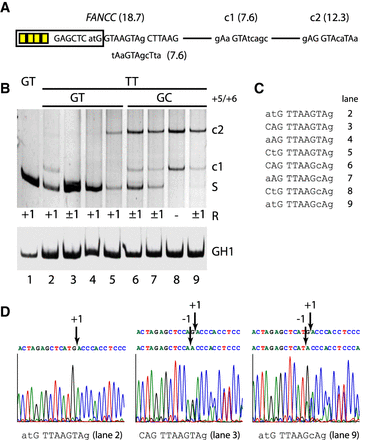
Accuracy of noncanonical TT splicing. (A) Schematic of the splicing reporter containing the human FANCC exon 2 splice site and two weak cryptic splice sites (c1/c2) in the downstream intron. Exonic SRSF7 enhancer sites are indicated by yellow boxes. (B) Different splicing positions (R) obtained by variations of the TT splice site sequence. RT-PCR analysis was performed as described before. All experiments were performed in triplicate (Supplemental Fig. S6). (C) TT splice site sequences from B, lanes 2–9. The human pathogenic FANCC exon 2 c.165 + 1G > T splice site is shown in lane 2. (D) Exemplary splicing positions mapped by sequencing of the extracted RT-PCR products.











