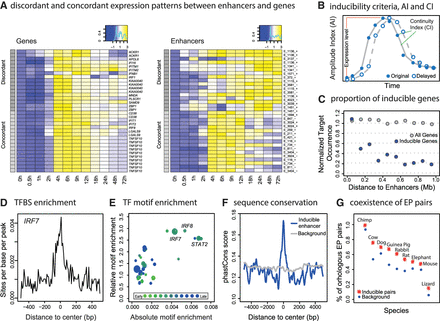
Prediction of virus-inducible enhancer-promoter (EP) pairs and validation of their interactions. (A) Heat map of discordant and concordant expression pairs of target genes (left panel) and their enhancers (right panel). Rows are matched. Expression levels were normalized as log2 fold changes relative to 0 h. Gray scales represent discordant (dark gray) and concordant (light gray) groups. (B) Diagram describes the identification of inducible enhancers and genes with two indices: the continuity index (CI) and the amplitude index (AI). (C) Number of inducible genes as a function of EP distance is analyzed. Inducible genes were counted within each 100-kb bin of inducible enhancers (blue points). As a control, the number of all genes in each bin was calculated (gray points). Each group was normalized by the maximum count for the sake of comparison. (D) Enrichment of TF IRF7 motif in the 1-kb TSS-flanking region of inducible enhancers is shown. (E) Motif enrichment of inducible enhancers. x-axis (absolute enrichment) is the maximum sites per base per peak (SBP). y-axis (relative enrichment) is the SBP ratio between the center (−100 to 100 bp) and rest (−500 to −100 bp and 100 to 500 bp) of the flanking regions. Point sizes indicate the GRO-seq RPKM fold changes of TFs. Colors indicate the first time point when the TF reaches half induction. (F) Average phastCons conservation scores of primates across inducible enhancer regions are shown, relative to randomly selected background. (G) Percentage of inducible human EP pairs that co-exist in other species is shown.











