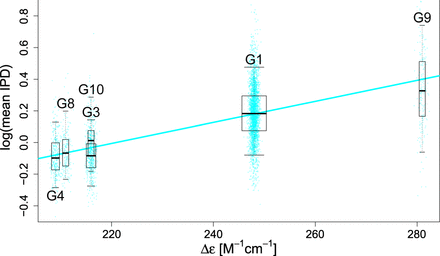
Relationship between G-quadruplex stability and polymerization kinetics. For the 10 most common G-quadruplex motif types (G1 through G10, in order), we measured circular dichroism (delta epsilon) and light absorption (melting temperature [Tm]) and computed average IPD values for each of thousands of motif occurrences in the genome (Supplemental Table S5). For intramolecular G4s, average IPDs were regressed on delta epsilon (R2 = 32.3%). (Box plot whiskers) 5th and 95th quantiles; (box plot width) proportional to the square root of the sample size for each motif; (points) individual occurrences used in the regressions, with horizontal jittering for visualization (results for delta epsilon in intermolecular G4s and for Tm are shown in Supplemental Fig. S15).











