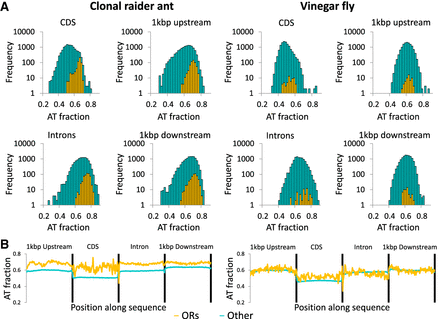
Nucleotide composition of odorant receptors (ORs) vs. other genes in the clonal raider ant and the vinegar fly (Drosophila melanogaster). (A) Frequency distribution of AT content for coding sequence (CDS), introns, downstream, and upstream sequence of OR genes (orange) and other genes (cyan). Y-axes are log-scaled to allow visualization of fly ORs, which make up a small proportion of fly genes. (B) Mean nucleotide composition along the CDS, intronic, and flanking sequences of OR genes or other genes in the clonal raider ant (left) and the vinegar fly (right). These data show that clonal raider ant ORs are AT-rich relative to most other genes, while fly ORs possess typical AT content relative to other genes.











