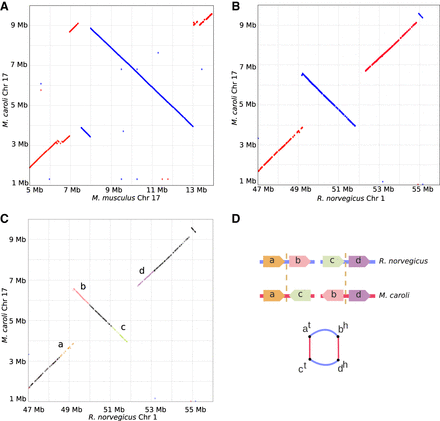
Large inversion detected in M. caroli Chr 17. Genome dot-plots of a Chromosome 17 region of M. caroli against M. musculus (A) and R. norvegicus (B) showing a cluster of structural variations of size ∼5 Mb. M. pahari chromosomes shows the same genomic structure. The inversion breakpoints are supported by the optical maps in both the M. caroli and M. pahari genomes. M. musculus and R. norvegicus references share one breakpoint of two different inversions. (C,D) An illustration of the chimera detection algorithm. The same inversion with highlighted synteny blocks (C) forms an alternating cycle of length four in the breakpoint graph (D). Thus, the Ragout 2 chimera detection algorithm classified these red edges as genomic (not artificial) and included the corresponding adjacencies into the assembled chromosomes.











