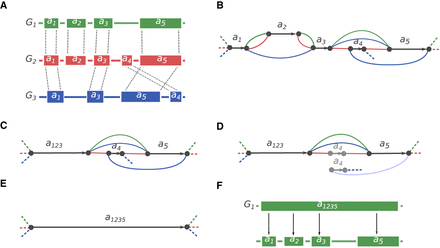
Synteny blocks reconstruction algorithm. (A) Three genomes—G1, G2, G3—are encoded in the alphabet of local sequence alignment blocks. |a4| < |a1| < |a3| < |a2| < |a5|. Alignment is performed using the Cactus multiple whole-genome aligner. (B) The A-Bruijn graph constructed with minBlock = |a4|. Black edges correspond to the alignment blocks, and the colored edges connect the adjacent alignment blocks in the corresponding genome. Dashed edges denote the start/end of each genome. Because the minBlock parameter is equal to the size of the smallest block, all blocks were included in the graph. (C) The A-Bruijn graph after bubble simplification and collapsing unbranching paths. a123 now represents the new merged block (from a1, a2, a3). (D) A next iteration of the A-Bruijn graph with |a4| < minBlock ≤ |a1|, which eliminates block a4. (E) a123 and a5 are merged into a larger block a1235. (F) The hierarchical representation of synteny blocks: The larger block a1235 from the genome G1 can be decomposed into smaller blocks a1, a2, a3, and a5.











