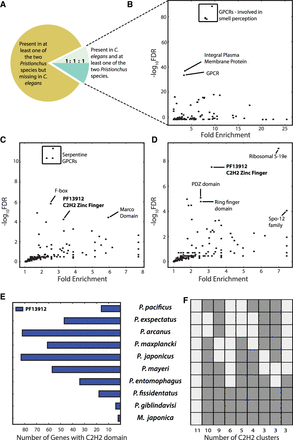
P. pacificus–specific loss. (A) Majority of gene families that have been lost in P. pacificus, but have at least one gene from either P. exspectatus or P. arcanus, do not have any orthologous genes in C. elegans. (B) Gene Ontology analysis of C. elegans orthologs of P. exspectatus and P. arcanus genes whose counterparts were lost in P. pacificus shows an enrichment of G-protein coupled receptors. (C,D) Overrepresentation of protein domains among genes that have been lost in P. pacificus based on orthologs from P. exspectatus (C) and P. arcanus (D). The C2H2-type zinc finger domain (PF13912) shows a consistently significant enrichment in both species. (E) The number of genes with C2H2 domains across all 10 species indicates an expansion of this domain in the Pristionchus lineage. (F) The nine most abundant species distribution patterns in orthologous clusters containing a C2H2 domain show additional expansions and contractions.











