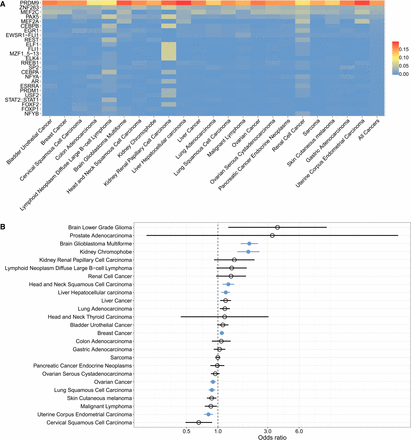
Structural variant (SV) breakpoints are enriched at sites of PRDM9 binding and activity. (A) Proportion of SV breakpoint sequences (SVBSs) significantly matching the motifs recognized by proteins in the JASPAR database, in cancer samples expressing PRDM9. We focused on sequences located within 100 bp of SV breakpoints, which is the mean distance separating DSBs and PRDM9 binding sites in meiosis (Baker et al. 2014). Each row shows the proportion of SVBSs matching the given motif per cancer type. The far-right column shows the proportion of SVBSs matching each motif across all cancer types. Supplemental Figure S7 shows the robustness of these results to significance thresholds used (Methods). (B) Enrichment for SV breakpoints at sites of highly recombining regions (HRRs) in samples expressing PRDM9, as shown by odds ratios >1 for each cancer type. Significant cancer types are shown in blue, as determined using Fisher's exact test (P < 0.05 with Bonferroni correction for the number of cancer types tested). Brain glioblastoma multiforme, kidney chromophobe, head and neck squamous cell carcinoma, liver hepatocellular carcinoma, and breast cancer samples all exhibited significant associations between PRDM9 expression and the colocalization of SV breakpoints and meiotic recombination hotspots. Ovarian cancer, lung squamous cell carcinoma, and uterine corpus endometrial carcinoma showed odds ratios <1, indicating significant enrichment for SV breakpoints in non-HRRs in samples expressing PRDM9.











