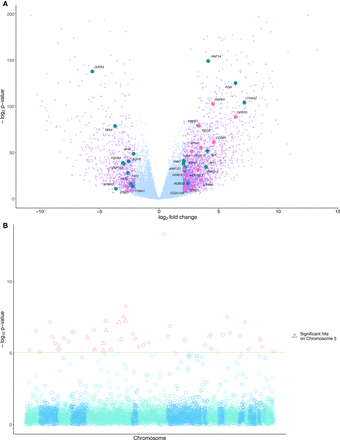
Differentially expressed genes and recurrently mutated regions are associated with PRDM9 expression in cancer. (A) A total of 3114 genes were differentially expressed (in purple) above a threshold of 2 log2 fold change and a P-value adjusted for multiple testing using the Benjamini-Hochberg procedure (FDR < 0.05) between groups of cancers partitioned on PRDM9 expression. Of these genes, 2224 were overexpressed in cancers expressing PRDM9, and 890 were underexpressed. Twenty-two known cancer driver genes (Gonzalez-Perez et al. 2013) were differentially expressed in tumors expressing PRDM9 (in teal), and 13 had functions related to meiosis (in pink). (B) P-values from the associations between PRDM9 expression levels and recurrently mutated regions. The red line shows P < 0.05, with a Bonferroni correction for multiple testing for the number of tested regions (n = 1,507,106). Forty-nine recurrently mutated regions were significantly associated with aberrant PRDM9 expression across cancers. Highlighted with pink triangles are loci significantly associated with aberrant expression located on Chromosome 5, where the PRDM9 locus is located.











