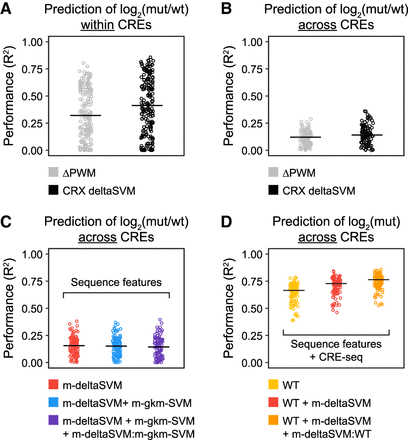
Accounting for baseline CRE activity improves the prediction of variant effects. (A) Performance (R2) of simple linear regression predicting the effect of individual substitutions from changes in PWM scores or CRX ChIP-seq deltaSVM scores, fitting separate models for each CRE. (B) Same as in A, except fitting a single model for all CREs. (C) Performance of multiple linear regression predicting the effect of individual substitutions using deltaSVM scores from multiple data sets (m-deltaSVM), m-deltaSVM and the corresponding gkm-SVM scores from multiple data sets (m-gkm-SVM), or m-deltaSVM scores and m-gkm-SVM scores including all pairwise interactions. (D) Performance of multiple linear regression predicting mutant expression using wild-type (WT) expression, WT expression and m-deltaSVM scores, or WT expression, m-deltaSVM scores, and interactions between WT expression and deltaSVM scores. In A, individual points represent the performance of models fit for different CREs (n = 195). In B–D, individual points represent the performance of models estimated from different folds of repeated 10-fold cross-validation (n = 100).











