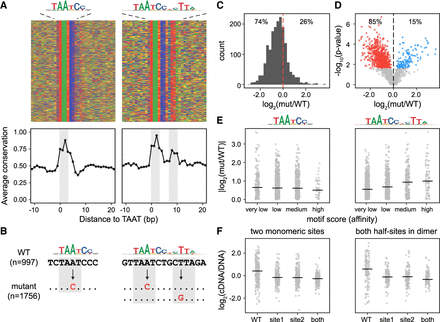
Dimeric CRX sites have higher activity than monomeric CRX sites. (A, top) Heatmaps of nucleotide content in a 30-bp window centered on monomeric or dimeric CRX binding sites. Rows correspond to distinct TF binding sites, columns correspond to distinct positions, and tiles are colored by nucleotide identity. (Bottom) Average conservation (100-way vertebrate phyloP scores) at each position. Positions 0–3 (and 7–10 for dimeric TF binding sites) correspond to TAAT cores (gray boxes). (B) Schematic of experimental approach. The effects of single–base pair substitutions (TAAT to TACT) in 1756 CRX binding sites within CRX ChIP-seq peaks were quantified by CRE-seq. (C) Distribution of mutation effects (log2 fold change). (D) Volcano plot of mutation effects. Among mutations that significantly change activity (FDR < 0.05), 85% decrease activity and 15% increase activity. (Red) Significant decrease in activity; (blue) significant increase in activity; and (gray) change in activity not significant. (E) Absolute effect size versus monomeric or dimeric PWM score (binned by match P-value). (F, left) Activity distributions of CREs with two monomeric CRX binding sites when neither, one, or both are mutated. (Right) Activity distributions of CREs with dimeric CRX binding sites when neither, one, or both half-sites are mutated.











