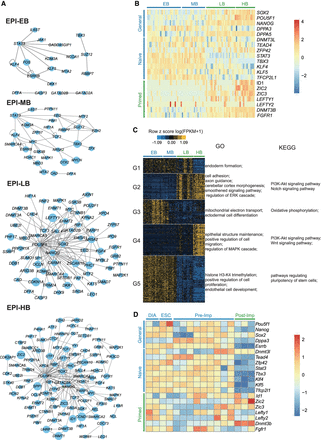
Distinct pluripotent states exist in rhesus monkey pre-implantation embryos. (A) PluriNetWork analysis of EPI cells from four blastocyst stages revealed a transition of network topology from mid- to late-blastocyst stage. (B) Expression pattern of marker genes for naive pluripotency, primed pluripotency, and general pluripotency in EPI cells of EB, MB, LB, and HB in rhesus monkey. (C) BICSKmeans clustering of differentially expressed genes (DEGs) (FDR < 0.01) in EPI cells from EB through to HB stages. The GO term and KEGG pathway enrichment of each group of DEGs are shown. In A–C, data were from 90 batch 2 EPI cells (EB, 29 cells; MB, 20 cells; LB, 24 cells; and HB, 17 cells). (D) Expression pattern of marker genes for naive pluripotency, primed pluripotency, and general pluripotency in EPI cells of pre-implantation and post-implantation embryos in mouse. Analysis was performed on the archival data from Boroviak et al. (2015).











