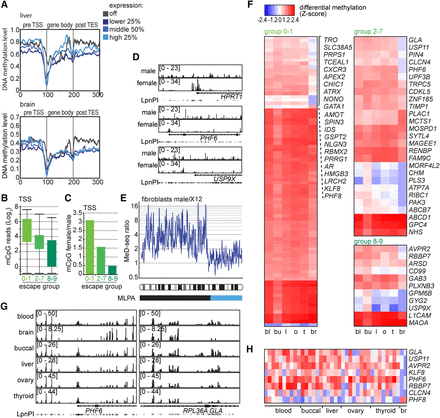
Organ and patient-specific escape of XCI. (A) Gene density plots showing DNA methylation profiles for genes of four different expression groups for liver (top) and brain (bottom). DNA methylation level is shown in 100 bins before the TSS, the gene body, and behind the transcriptional stop. (B) TSS-specific MeD-seq reads (Log2) for genes subject to XCI (group 0–1), partially escaping XCI (groups 2–7), and nearly completely escaping XCI (groups 8–9) in liver. (C) Bar graph showing the ratio of TSS methylation of X-linked genes, subject to XCI (group 0–1), partially escaping XCI (groups 2–7), and nearly completely escaping XCI (groups 8–9), in female versus male fibroblasts. (D) MeD-seq tracks displaying DNA methylation in genes subject to XCI (HPRT, top), partially escaping XCI (PHF6, middle), and escaping XCI (USP9X, bottom). (E) Ratio of TSS read counts for female X12 deletion and male control fibroblasts along the X Chromosome. The sliding average of 10 TSSs is shown, and the exact break point determined by MLPA analysis is shown below (light blue). (F) Inter-organ DMRs were called and displayed according to the escape group. TSS DMRs are shown that were significant for all tissues by Bonferroni testing, colored according to the Z-score. (G) MeD-seq tracks of TSS-specific DMRs for PHF6 and GLA that show hypomethylation in brain. (H) Z-scores for TSSs of genes selected based on a brain-specific methylation pattern of 0–1, 2–7, and 8–9 genes shown in organs of individual patients.











