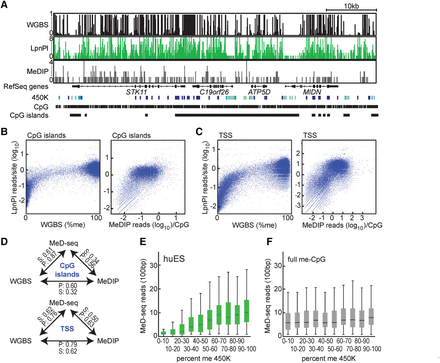
MeD-seq versus WGBS, MeDIP, and the Infinium 450K technology. (A) MeD-seq, WGBS, and MeDIP DNA methylation profiles and Infinium 450K scores for a representative locus on Chromosome 19, obtained with human ES cell DNA. MeD-seq and MeDIP read count or relative DNA methylation level (0–1) is shown for all CpG sites. Infinium 450K scores: (dark blue) high CpG methylation; (light blue) intermediate DNA methylation; (green) low DNA methylation (based on B-value scores). (B) Correlation plots for MeD-seq:WGBS (left) and MeD-seq:MeDIP (right) comparison of CpG islands. (C) As in B, but now for TSS. (D) Overview of Pearson (P) and Spearman (S) correlation scores for different comparisons. (E) MeD-seq reads in a 100-bp window plotted against Infinium 450K scores binned in 10 different groups with increasing methylation scores for ES cell DNA. (F) As in B, but now with MeD-seq reads obtained from 100% methylated control DNA using 450K bins obtained with ES cell DNA.











