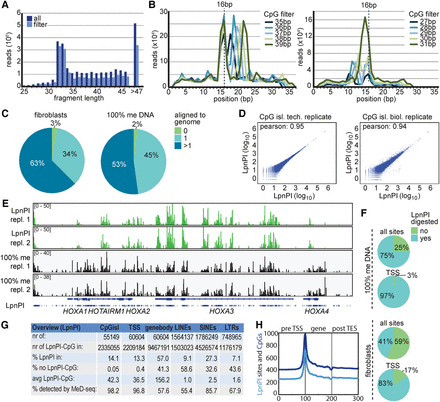
Detection of DNA methylation genome wide. (A) Histogram showing sequencing read size of all reads (dark blue) and reads that passed the LpnPI filter (light blue) obtained with MeD-seq applied on human fibroblast DNA. (B) Sequencing reads larger (top) or shorter (bottom) than 32–33 bp shown in F display two LpnPI recognition sequences 16–17 bp from the end. (C) Alignment of LpnPI-filtered MeD-seq reads obtained using fibroblast DNA (left) and 100% methylated DNA (right). (D) Pearson correlation analysis of MeD-seq read counts of CpG islands for technical (left) and biological (right) replicates. (E) MeD-seq DNA methylation profiles of the HOXA cluster obtained with fibroblast DNA (top two panels) and 100% methylated control DNA (bottom two panels). (F) Ratio of digested LpnPI sites in CpG islands and TSSs for 100% methylated control DNA (top) and fibroblast DNA (bottom). (G) Overview and characteristics of LpnPI sites in CpG islands, TSSs (2 kb), gene bodies, and repetitive elements genome wide. Detection percentages are based on 100% methylated DNA. (H) Gene density plots showing distribution of LpnPI sites and CpG dinucleotides shown in 100 bins before the TSS, in the gene body, and behind the transcription stop.











