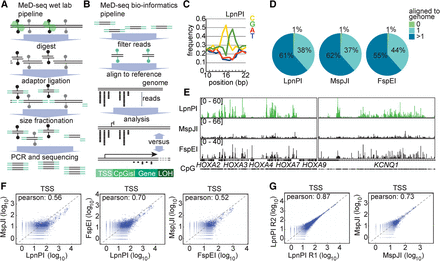
MeD-seq wet laboratory and bioinformatics platform. (A) Genomic DNA is digested with LpnPI, followed by DNA repair, adaptor ligation to 32-bp fragments, size fractionation, amplification, and sequencing. (B) Sequencing reads are trimmed, filtered based on the CpG sequence, and aligned to the genome. (C) Nucleotide frequency plotted against the position in the sequencing reads, showing enrichment of CG nucleotides around 16–17 bp from the start. (D) Alignment of LpnPI, MspJI, and FspEI CpG-filtered MeD-seq reads obtained using fibroblast DNA. (E) Sequencing read profiles of the HOXA and KCNQ1 loci obtained with fibroblast DNA using LpnPI, MspJI, and FspEI. (F) Pearson correlation analysis of MeD-seq read counts of TSSs for LpnPI-MspJI (left), LpnPI-FspEI (middle), and FspEI-MspJI (right) comparisons. (G) Pearson correlation analysis of read counts of TSSs for technical replicates digested with LpnPI (left) and MspJI (right).











