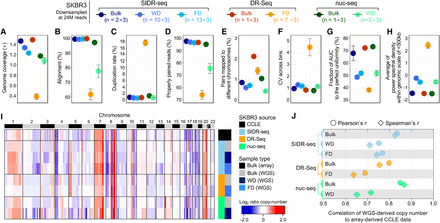
Comparison of single-cell genome sequencing methods: SIDR-seq, DR-Seq, and nuc-seq. The number of sequencing reads in each sample was set to 24 million in triplicate by randomly down-sampling from all available reads. (A–E) Summary of sequencing metrics. (A) Genome sequencing depth of coverage. The plots display fractions of sequencing reads properly aligned to the human reference genome (B), duplicated (C), properly paired (D), and with their paired reads mapped to different chromosomes (E). (F) Bin-to-bin variabilities in genomic DNA read counts. (G) Comparison of coverage uniformities measured by Lorenz curves. The fractions of area under the curve were calculated, averaged for each group, and plotted. (H) Comparison of coverage uniformities measured by power spectral analysis. Power spectral densities of read distributions were obtained and averaged across frequencies >1/500 kb. (I) Heatmap of genome-wide copy-number profiles in bulk and single cells from SKBR3 cells by binning of 1-Mb genomic scale. Copy-number profiles from genome sequencing were compared to the CCLE data profiled using SNP array (at the top of the heatmap). (J) Correlation of copy numbers between data sets from each method and CCLE data set. Pearson's and Spearman's correlation coefficients were plotted against the x-axis. (FD) DNA fractionated from single cells by SIDR; (WD) genomic DNA from the whole-cell lysates of single cells.











