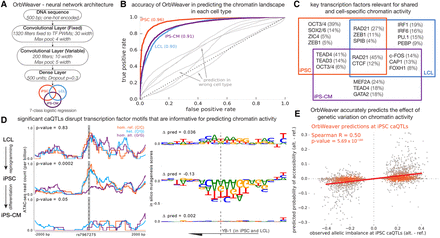
Predicting chromatin activity from sequence using deep neural networks. (A) OrbWeaver is a four-layered neural network where the parameters of the first convolutional layer are fixed to known position weight matrices of human transcription factors. The activation function used in each of the convolutional and dense layers is the Rectified Linear Unit (ReLU). (B) The OrbWeaver model for one cell type poorly predicts open chromatin in other cell types (gray), highlighting that the model captures cell-type–specific regulatory elements. (C) Transcription factors important for each locus were identified using DeepLIFT scores; this panel illustrates the top key TFs for each of the seven categories of chromatin activity and the fraction of loci explained by them. (D) An example of a locus that is open in iPSCs and LCLs but was identified to be an iPSC-specific caQTL. The subpanels on the left show the raw ATAC-seq signal in each cell type stratified by genotype of the most significant SNP of the iPSC caQTL. The subpanels on the right show the marginal change in OrbWeaver predictions due to mutating the reference base at each position to an alternate base. The sequence shown corresponds to the shaded portion on the left subpanels, and the reported Δpred values correspond to the change between alleles of the most significant SNP. The TF important for this locus as identified by DeepLIFT is YB-1, a factor highly expressed in all three cell types. (E) Scatter plot comparing the observed allelic imbalance at iPSC caQTLs, estimated by WASP, and the predicted difference in median chromatin activity between haplotypes tagged by the two alleles of the causal SNP. Note that the OrbWeaver model was learned using the reference genome sequence alone and had no information regarding genetic variation in the population when learning the model parameters.











