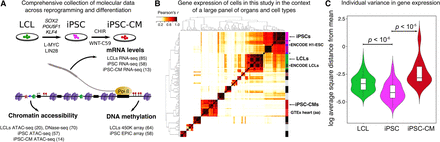
Systematic measurements of molecular phenotypes across reprogramming and differentiation. (A) Summary of data collection. (B) Correlation matrix of gene expression from our samples and samples from ENCODE (*) and GTEx. Our LCL samples cluster most closely with LCLs samples from ENCODE, while our iPSCs and iPSC-CM lines cluster most closely with H1-ESC (ENCODE) and heart (GTEx), respectively. Dark purple: GTEx bone marrow. (C) Violin plots representing per individual log2 of the average square distance from the mean (Supplemental Materials) for iPSC, LCL, and iPSC-CM gene expression levels. Plots for chromatin accessibility and DNA methylation levels are shown in Supplemental Figure S7.











