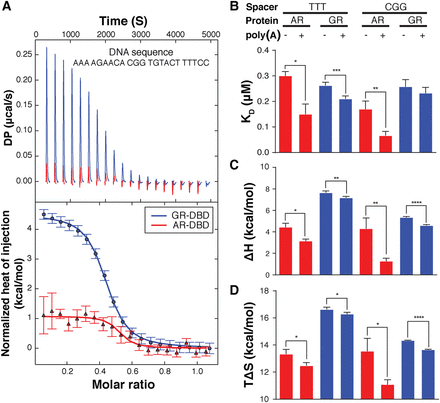
ITC analysis reveals distinct DNA-binding thermodynamics between AR- and GR-DBD. (A) The raw heat titration signals (top) and normalized heat of injection profiles (bottom) of AR- and GR-DBD bound to a given DNA sequence. Standard errors are estimated by NITPIC (Brautigam et al. 2016). (B) The KD of AR-DBD and GR-DBD for four sets of sequences fit from the ITC data. AR-DBD affinity is increased with flanking As and an optimal spacer, whereas GR is insensitive. (C) Enthalpy, ΔH, is calculated from the heat of binding for each DNA sequence. Flanking sequences decrease ΔH for AR-DBD, enhancing affinity. Smaller indicates a greater contribution to affinity. (D) Entropy, ΔS, is calculated from the KD (thus ΔG) and ΔH. GR-DBD affinity is more entropically driven. Larger indicates a greater contribution to affinity. (*) P-value ≤0.05, (**) P-value ≤0.01, (***) P-value ≤0.001, (****) P-value ≤0.0001, two-sided t-test. Error bars, SD represent the standard deviation from at least three experiments.











