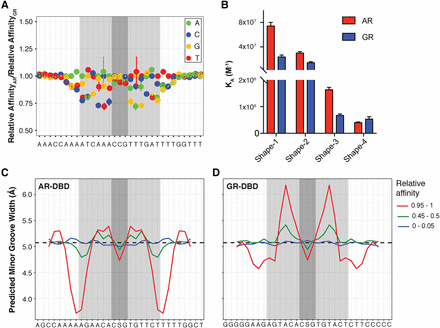
Difference in DNA shape readout between AR and GR. (A) Difference in ΔΔG/RT values between AR and GR at each nucleotide position, normalized by their mean across all four bases. (B) Quantitative EMSA was used to measure the affinities of AR- and GR-DBDs for four sequences that maximally favor AR (Shape-1) and GR (Shape-4) and test the importance of half-site minor groove width (Shape-3 and -4). Each measurement was performed at least three times. Error bars, SEM. Average minor groove width profile for the top/middle/bottom 5% of sequences in terms of affinity for AR (C) and for GR (D).











