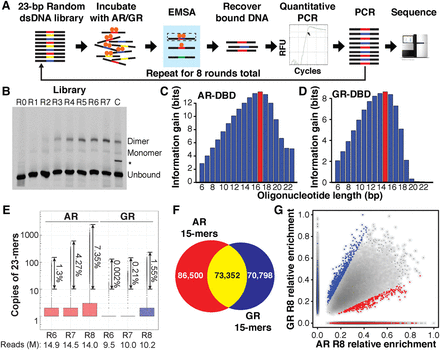
SELEX-seq reveals differences in AR- and GR-DBD (DNA-binding domain) DNA-binding specificity. (A) SELEX-seq. A 70-bp dsDNA library with 23-bp randomized region was incubated with the DBD of AR or GR and separated into monomer and dimer species by EMSA. Dimer-bound DNA was recovered, quantified by qPCR, amplified as the library for the next round, and repeated for eight rounds. Each round of library, including the initial dsDNA library, was sequenced. (B) EMSA gel showing the enrichment of dimer-bound sequences after each round of selection for GR-DBD. The intensity of the shifted band plateaus after round 7. A high-affinity palindromic sequence served as a control to locate the dimer band. (*) An artifact during the synthesis of control sequence but not observed in the SELEX library. (C,D) Information gain, or Kullback-Leibler divergence, from R0 to R8, as a function of oligonucleotide length. (E) Boxplot showing the multiplicity of unique 23-mers in each of the last three rounds of SELEX-seq selection for AR and GR. Even for the most highly selected library (AR R8) fewer than 10% of all reads have 10 copies or more, indicating that the libraries are not overselected. (F) Venn diagram showing the overlap of sequences for AR- and GR-DBD with at least 100 sequencing counts. (G) Scatterplot of sequences that were commonly bound (yellow from F) by AR- and GR-DBD.











